Types of Genomic Copy Number Variations
Classification of CNVs based on CN level and genomic size
Genomic copy number variations can affect the production of proteins through their modification of the "dosage" (i.e. number of alleles) of genes covered by the CNV as well as through the (in)complete disruption of coding or regulatory elements. The supposed effects depend on the magnitude of change, i.e. the number of gained or lost copies. Especially in cancer cells the "normal" allele count for the given genomic region has to be considered: Cancer genomes may have undergone general anaeuplodization events with preceding and/or followed by regional CNV events.
Copy number changes can be expressed as absolute (total allele count) and relative (e.g. uncalibrated CN measurements such as log2 ratio) values. Empirically - and to overcome a number of issues of exact CN count calibrations - a classification into a number CN types has become "standard operating procedure" with details differing to some extent. This post attempts to summarize some previous annotation practices and emerging standards for CNV annotation, especially with respect to somatic CNVs and cancer genomics.
Continue reading1000 Genomes CNV reference data
An API for the WGS-based reference dataset of CNVs in the 1000 Genomes samples
In October 2020, the AWS for Industries group published a re-analysis of 3202 deep WGS sequencing datasets from the 1000 Genomes Project (1kGP), freely available for download by interested researchers and including the copy number variations (CNV) inferred for these genomes. Based on a set of well defined samples, this data can be considered a "high-quality resource for germline CNV data", for disease and population analysis projects. Now, this data is being made available though the Progenetix resource's API.
1000 Genomes Germline CNVs on Progenetix
LINK: progenetix.org/progenetix-cohorts/oneKgenomes/¶
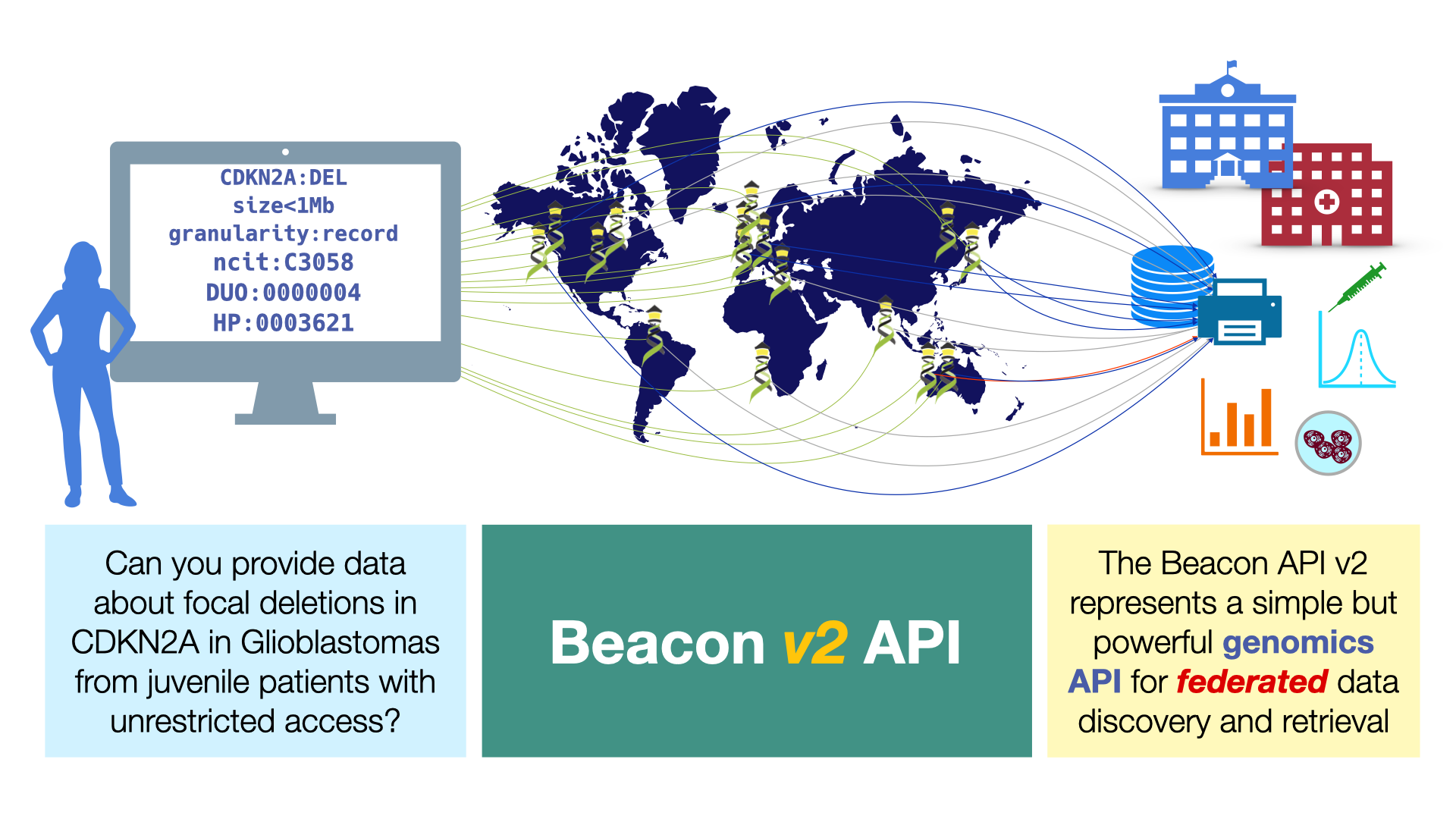
GA4GH Approves Beacon v2
A major step towards federated analysis of biomedical genomics data
![]() At its Spring 2022 GA4GH Connect stakeholder meeting in Montreal (and online...) the
steering committee of the Global Alliance for Genomics and Health (GA4GH)
approved the major "v2" update of the Beacon protocol as official GA4GH standard.
At its Spring 2022 GA4GH Connect stakeholder meeting in Montreal (and online...) the
steering committee of the Global Alliance for Genomics and Health (GA4GH)
approved the major "v2" update of the Beacon protocol as official GA4GH standard.
CNVs in Prenatal Tests & Maternal Malignancies
Publication indicating rare CNV signatures from a nationwide Dutch screening program
In a new publication in the Journal of Clinical Oncology CJ Heesterbeek, SM Aukema and the co-authors from the Dutch NIPT Consortium report about the incidence and diagnostic significance of incidential detection om maternal copy number variations in a large screening program aimed at detecting chromosomal imbalances in embryos, for a prediction of developmental abnormalities.
Citation
Noninvasive Prenatal Test Results Indicative of Maternal Malignancies: A Nationwide Genetic and Clinical Follow-Up Study.
Heesterbeek CJ, Aukema SM, Galjaard RH, Boon EMJ, Srebniak MI, Bouman K, Faas BHW, Govaerts LCP, Hoffer MJV, den Hollander NS, Lichtenbelt KD, van Maarle MC, van Prooyen Schuurman L, van Rij MC, Schuring-Blom GH, Stevens SJC, Tan-Sindhunata G, Zamani Esteki M, de Die-Smulders CEM, Tjan-Heijnen VCG, Henneman L, Sistermans EA, Macville MVE, Dutch NIPT Consortium.
J Clin Oncol PMID:35394817 | JCO
GA4GH implementation study - Making ontologies work
Diagnoses, phenotypes, species assignment and other “biocharacteristics” represented as “OntologyTerm” objects
Historical Post
This post appeared first (on WordPress hosted genome.blog) on 2017-04-13 and has been slightly edited to provide the evolution of the concepts. The post is a reminder of the history of some data structures now implemented throughout GA4GH aligned projects.
While developing the (metadata-)schema for the Global Alliance for Genomics and Health, the Metadata Task Team decided (after, well, many hours of telecons) to use an object based model with a limited number of attributes. In place of specific, named attributes, emphasis is being put on the use of ontologies, which, in principle, provide their own scoping. Diagnoses, phenotypes, species assignment and other “biocharacteristics” were to be represented in as “OntologyTerm” objects, using schema-external ontology services such as EBI’s OLS.
Continue readingThe ELIXIR Human Copy Number Variations Community
Whitepaper from the ELIXIR hCNV community
In this whitepaper, members of the newly founded ELIXIR hCNV community describe the reasons for and the expected trajectory of the newly assembled interest group, supported as one of the ELIXIR communities.
Citation
The ELIXIR Human Copy Number Variations Community: building bioinformatics infrastructure for research
Salgado D1, Armean IM, Baudis M, Beltran S, Capella-Gutierrez S, Carvalho-Silva D, Dominguez Del Angel V, Dopazo J, Furlong LI, Gao B, Garcia L, Gerloff D, Gut I, Gyenesei A, Habermann N, Hancock JM, Hanauer M, Hovig E, Johansson LF, Keane T, Korbel J, Lauer KB, Laurie S, Leskosek B, Lloyd D, Marques-Bonet T, Mei H, Monostory K, Pinero J, Poterlowicz K, Rath A, Samarakoon P, Sanz F, Saunders G, Sie D, Swertz MA, Tsukanov K, Valencia A, Vidak M, Yenyxe Gonzalez C, Ylstra B, Béroud C
Markers, Markers Everywhere - Will they Kill Me? Should I Care?
Reporting of SNVs identified in GWAS studies in news
This can be very brief: Since the arrival of genomic arrays aimed at mapping Single Nucleotide Polymorphisms (SNP), and later sequencing based variation profiling, there has been a deluge of (more or less) well designed studies to look for a statistical correlation between the occurrence of SNP variants at genomic loci, and (more or less) well defined traits or phenotypes.
Continue reading